Who's got the biggest workload? Analysis of chaperone abundance and workload
Brownridge P, Lawless C, Payapilly AB, Lanthaler K, Holman SW, Harman VM, Grant CM, Beynon RJ, Hubbard SJ. Quantitative analysis of chaperone network throughput in budding yeast (2012). Proteomics. 13(8) 1276-1291 [PUBMED][PDF]
The network of molecular chaperones mediates the folding and translocation of the many proteins encoded in the genome of eukaryotic organisms. It has been particularly well characterised in the budding yeast, Saccharomyces cerevisiae, where 63 known chaperones have been annotated and recent affinity purification and tandem mass spectrometry experiments have helped characterise the attendant network of chaperone targets to a high degree. In this study, we apply our QconCAT methodology to directly quantify the set of yeast chaperones in absolute terms (copies per cell) via selected reaction monitoring mass spectrometry. Firstly, we compare these to existing quantitative estimates of these yeast proteins, highlighting differences between approaches. Secondly, we cast the results into the context of the chaperone target network and show a distinct relationship between abundance of individual chaperones and their targets. This allows us to characterize the “throughput” of protein molecules passing through individual chaperones and their groups on a proteome-wide scale in a model eukaryote for the first time. The results demonstrate specialisations of the chaperone classes, which display different overall workloads, efficiencies, and preference for the subcellular localisation of their targets. The novel integration of the interactome data with quantification supports re-estimates of the level of protein throughout going through molecular chaperones. Additionally, we show that chaperones mediate the folding of the majority of protein molecules (~62% of the total protein flux in the cell), although chaperones target fewer than 40% of annotated proteins, highlighting their importance.
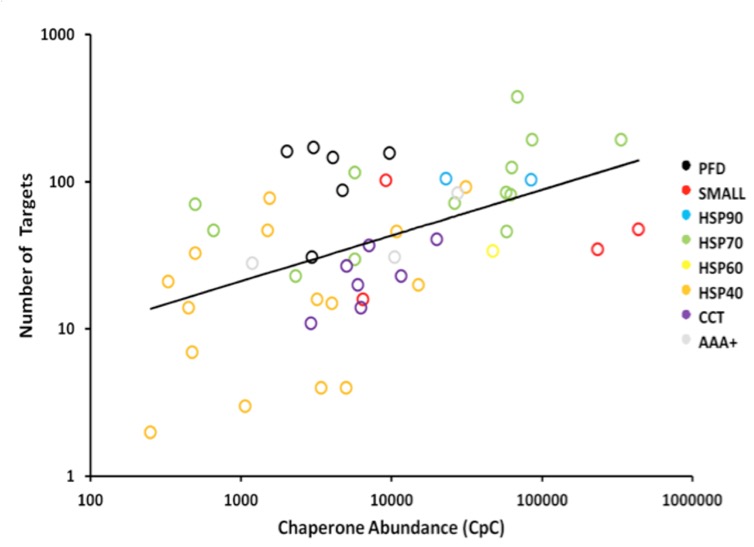
The network of molecular chaperones mediates the folding and translocation of the many proteins encoded in the genome of eukaryotic organisms. It has been particularly well characterised in the budding yeast, Saccharomyces cerevisiae, where 63 known chaperones have been annotated and recent affinity purification and tandem mass spectrometry experiments have helped characterise the attendant network of chaperone targets to a high degree. In this study, we apply our QconCAT methodology to directly quantify the set of yeast chaperones in absolute terms (copies per cell) via selected reaction monitoring mass spectrometry. Firstly, we compare these to existing quantitative estimates of these yeast proteins, highlighting differences between approaches. Secondly, we cast the results into the context of the chaperone target network and show a distinct relationship between abundance of individual chaperones and their targets. This allows us to characterize the “throughput” of protein molecules passing through individual chaperones and their groups on a proteome-wide scale in a model eukaryote for the first time. The results demonstrate specialisations of the chaperone classes, which display different overall workloads, efficiencies, and preference for the subcellular localisation of their targets. The novel integration of the interactome data with quantification supports re-estimates of the level of protein throughout going through molecular chaperones. Additionally, we show that chaperones mediate the folding of the majority of protein molecules (~62% of the total protein flux in the cell), although chaperones target fewer than 40% of annotated proteins, highlighting their importance.
